Cell and Gene Therapy: A Multifaceted Approach
02 July 2021 | Friday | Opinion
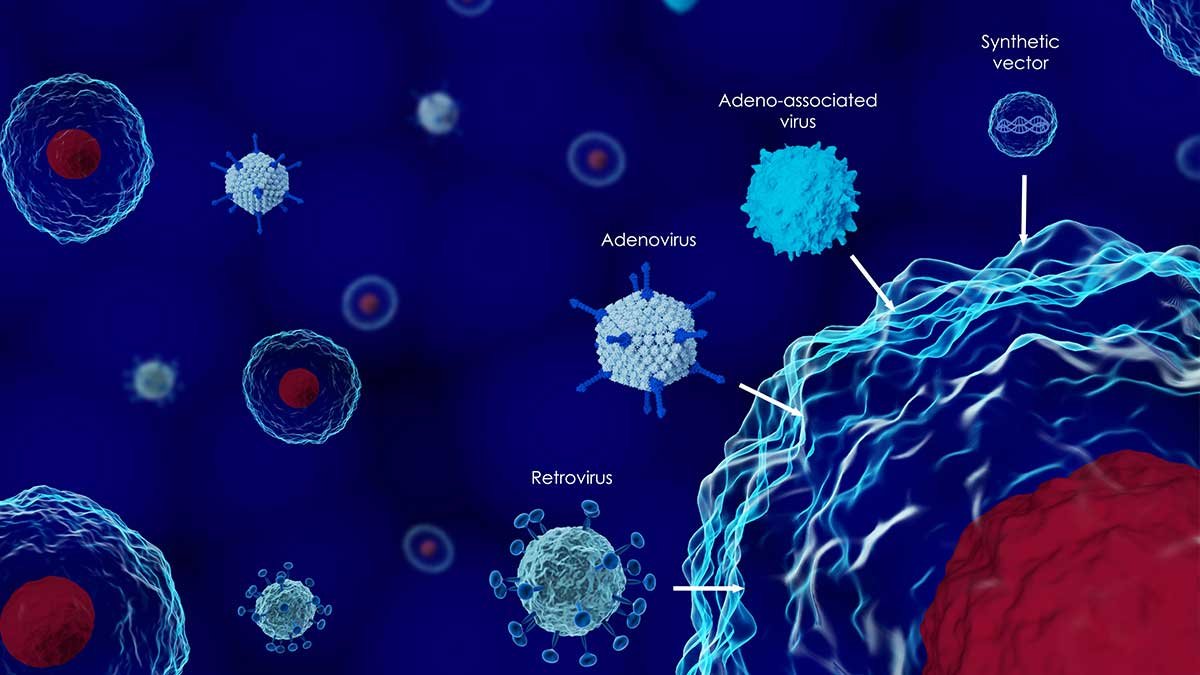
Source : Perkin Elmer
The active research in these areas not only encompasses laboratory bench work and imaging; it increasingly involves advanced computational methods in image processing, and machine learning to help tackle key problems.
Optimizing Cellular Delivery
While cell types are often characterized using immunological methods, it is now possible to classify cells solely based on their nuclear morphology without the use of antibodies. Scientists first generate detailed phenotypic profiles of stained nuclei from high-resolution microscopy images using texture (local intensity variations) and morphology parameters. Classification of nuclei then requires processing of data with machine learning algorithms.
Tools such as principal component analysis enable these algorithms to distinguish between up to six different types of simultaneously co-cultured cell types. The method is sensitive enough to distinguish between similar types of human cells with an over 90% accuracy rate. It has potential applications in distinguishing between healthy cells and intractable tumors.
Profiling and Characterizing the Delivery Vehicle
An important step in the development and quality control of the delivery vehicle is the quantitative assessment of its ability to transfect the right cells. Viral vectors rely on capsid proteins to convey cell type specificity (or tissue tropism), which are therefore the subject of intense research, and their identity is a crucial quality metric.
Speed and sensitivity are key considerations when characterizing capsid proteins, and both no-wash immunoassays and modern capillary electrophoresis methods offer relevant improvements over classic approaches. To measure viral transfection efficiency the plaque formation assay is well established. However, manual counting is time consuming and error prone especially at high virus titers. Automating this process greatly improves productivity.
Fast microscopic imaging of viral plaques allows the use of 96-well plates to dramatically improve throughput. Modern image analysis removes human error and enables detection of distinct plaques at high concentrations that are too difficult to count manually. It can also distinguish true viral plaques from cellular debris or contaminants.
Higher precision in viral assays allows for faster identification of novel viral capsid proteins required for tissue targeting and precise calculation of the viral titer. Image analysis of viral plaques distinguishes the delivery vehicle from cellular debris or contaminants, allowing for precise characterization of that vehicle’s mechanism of action. The delivery vehicle must be structurally and biochemically appropriate to the transgene, and its host is a key measure in quality control.
Delivery Vehicle Efficacy
While in vitro models are easy to use and provide valuable information, they are limited to cellular tropism and measuring the response of a cellular model. In vitro models are unable to accurately predict the biodistribution, viral gene expression, and degree of durability of the delivery vehicle. Therefore, small animal in vivo bioimaging plays an important role in both cell and gene therapies.
Within cellular therapies, in vivo bioimaging helps elucidate cellular populations and behaviors following transplantations. In gene therapies, where an adeno-associated virus (AAV) serotype is used with specific tissue tropism properties, only in vivo bioimaging can be used to study the targeting and efficacy of the therapy.
Successful gene therapy requires optimization of transgene delivery into target cells, precise characterization of the delivery vehicle, and careful monitoring of the delivery vehicle’s efficacy. This multi-pronged approach is supported by new image analysis technologies, machine learning, and mathematics. Cutting-edge methods of gene therapy have applications in cancer research, precise genetic targeting, and differentiation of cells for effective treatment.
PerkinElmer has developed a collection of application notes to support scientists in their analysis, characterization, and efficacy of the cell and gene therapy delivery vehicle.
Click Here to explore more
Most Read
- How Does GLP-1 Work?
- Innovations In Magnetic Resonance Imaging Introduced By United Imaging
- Management of Relapsed/Refractory Multiple Myeloma
- 2025 Drug Approvals, Decoded: What Every Biopharma Leader Needs to Know
- BioPharma Manufacturing Resilience: Lessons From Capacity Expansion and Supply Chain Resets from 2025
- APAC Biopharma Review 2025: Innovation, Investment, and Influence on the Global Stage
- Top 25 Biotech Innovations Redefining Health And Planet In 2025
- The New AI Gold Rush: Western Pharma’s Billion-Dollar Bet on Chinese Biotech
- Single-Use Systems Are Rewiring Biopharma Manufacturing
- The State of Biotech and Life Science Jobs in Asia Pacific – 2025
- Asia-Pacific Leads the Charge: Latest Global BioSupplier Technologies of 2025
- Invisible Threats, Visible Risks: How the Nitrosamine Crisis Reshaped Asia’s Pharmaceutical Quality Landscape
Bio Jobs
- Sanofi Turns The Page As Belén Garijo Steps In And Paul Hudson Steps Out
- Global Survey Reveals Nearly 40% of Employees Facing Fertility Challenges Consider Leaving Their Jobs
- BioMed X and AbbVie Begin Global Search for Bold Neuroscience Talent To Decode the Biology of Anhedonia
- Thermo Fisher Expands Bengaluru R&D Centre to Advance Antibody Innovation and Strengthen India’s Life Sciences Ecosystem
- Accord Plasma (Intas Group) Acquires Prothya Biosolutions to Expand Global Plasma Capabilities
- ACG Announces $200 Million Investment to Establish First U.S. Capsule Manufacturing Facility in Atlanta
- AstraZeneca Invests $4.5 Billion to Build Advanced Manufacturing Facility in Virginia, Expanding U.S. Medicine Production
News
Editor Picks